Limnic Habitats
Suspended Particulate Matter (SPM)
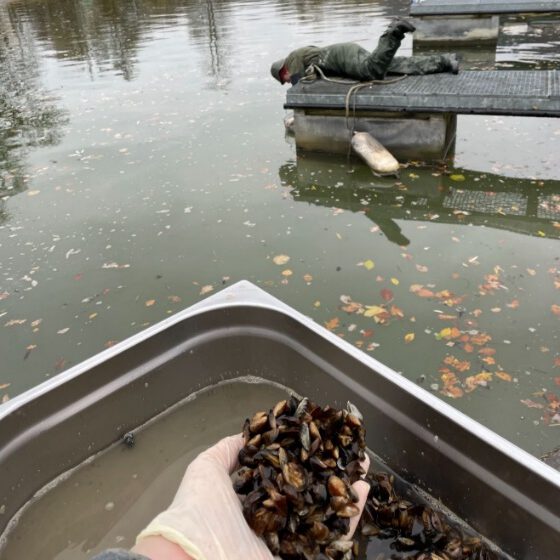
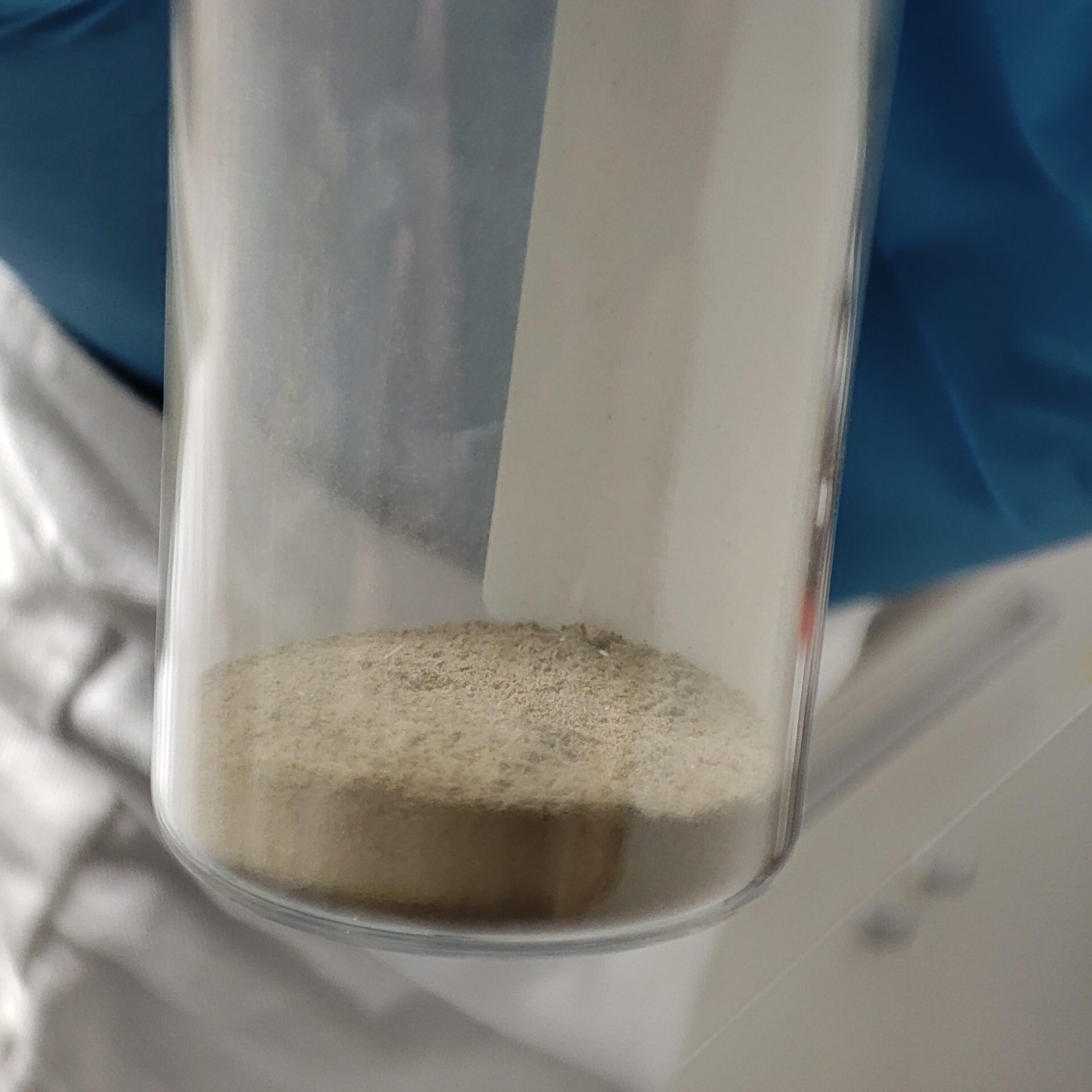
SPM samples have been taken monthly since 2005 at 12 sampling points on three major German rivers: the Rhine, the Elbe and the Danube. Samples are sieved on site and stored in liquid nitrogen. At the end of the year, the 12 monthly samples are mixed into an annual homogenate and stored. Monthly samples from the last few years are also available for comparison.
Using environmental DNA metabarcoding, the composition of the fish community as well as that of invertebrates and prokaryotes is analyzed over the time series of 2005-2020. Annual mixed samples for all 12 sampling points of the three rivers are being analyzed here (total: 312 SPM samples). Three to four markers are analyzed: COI (invertebrates), 12S (fish and other vertebrates), 16S (prokaryotes), and possibly an additional 18S (for broad eukaryote coverage). Correspondingly, three to four markers are generated for 312 measuring points (n=936 or 1248 data sets).
In addition to pure species detection and retrospective biodiversity monitoring, the data will be used to gain insights into the changes in species communities in the last 15 years. For example, where and when did non-native species
appear, how did they spread in the rivers over the years, when did native species disappear, and how did species on the Red List change in distribution?


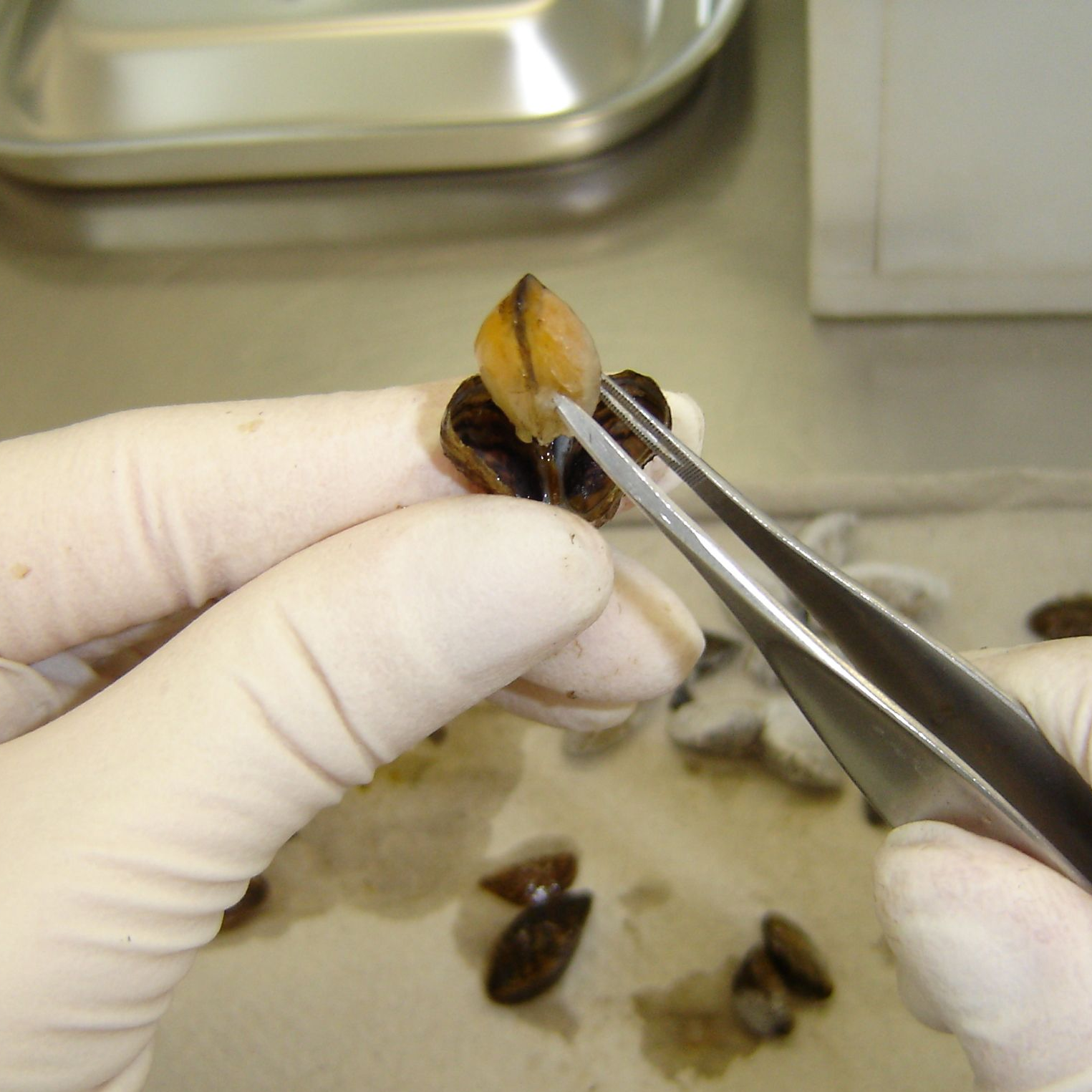
Zebra Mussels (Dreissena polymorpha)
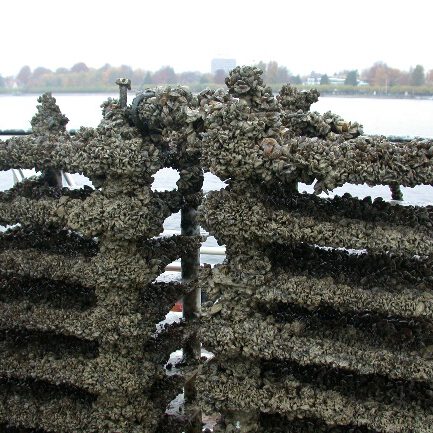
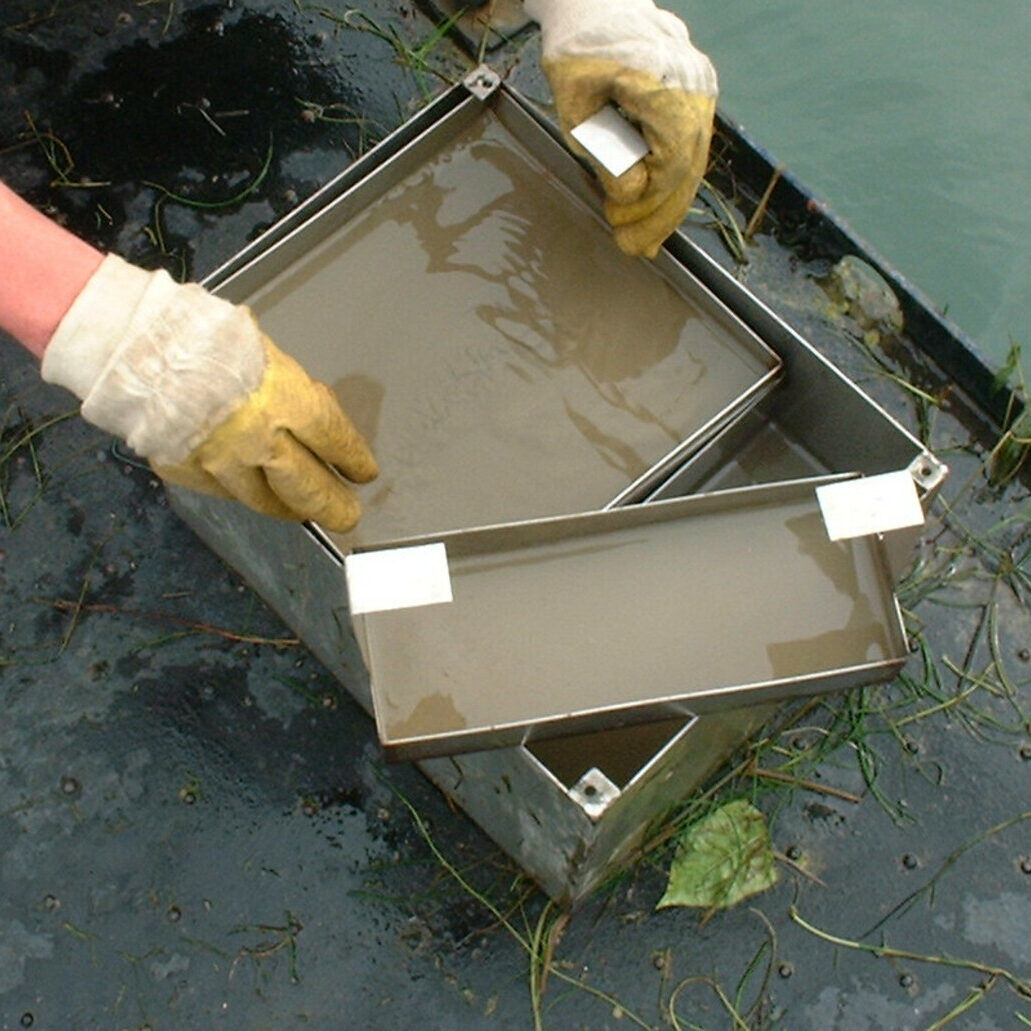
For the sampling of zebra mussels in the Danube, Elbe, Rhine and Saar as well as in the Belauer See, stacks of polyethylene plates are deployed in Lake Constance at the beginning of the year for mussels to colonize. After settlement, the plate stacks with the young mussels are removed to the sampling locations in the Danube, Elbe, Rhine, Saar and Belauer See. After a period of usually one year, the mussel samples are taken.
Sampling on the Danube: A specified minimum quantity of mussels is removed from each location after the end of the exposure period. If possible, the mussels are washed several times in the substrate water and coarse dirt particles are removed. Mussels with a length of less than 8 mm are excluded, as they fall through the mesh of the basket during the washing process. A total of 100 mussels are counted from the washed mussels in order to determine the ratio of zebra mussels (Dreissena polymorpha) to invasive quagga mussels (Dreissena rostriformes). The latter migrated into the German river system about 15 years ago and has already outcompeted the zebra mussel in many locations. For this reason, the quagga mussel is sampled here as a substitute species. At other locations, such as in Koblenz am Rhein, both species are sampled in parallel and sorted based on a key characteristic. Only zebra mussel samples from years in which contamination with the quagga mussel can be largely ruled out are used for the investigation of the time series within the TrendDNA project. Before being stored in liquid nitrogen at the sampling site, the mussels are spun dry as much as possible and blotted with paper towels. The mussels are then transferred to a stainless steel vessel, weighed and frozen over liquid nitrogen. In addition, 50 mussels are packed in a separate stainless steel container for biometric tests in the laboratory.
DNA metabarcoding is used to characterize the commensals and parasites associated with the mussel, as well as the planktonic food particles ingested by the mussel. This will give an overview of the surrounding community. 200 mussel homogenates are being analyzed using a three-marker multiplex assay that maps the zooplankton and phytoplankton communities around the mussel (Weber et al. 2021). The 200 homogenates come from 13 locations in five bodies of water (Saar, Rhine, Elbe, Danube, Belauer See) and cover a time series from 1990-2019.